Whole-Body [18F]FDG-PET/CT Imaging of Healthy Controls: Test/Retest Data for Systemic, Multi-Organ Analysis
![Whole-Body [18F]FDG-PET/CT Imaging of Healthy Controls: Test/Retest Data for Systemic, Multi-Organ Analysis Whole-Body [18F]FDG-PET/CT Imaging of Healthy Controls: Test/Retest Data for Systemic, Multi-Organ Analysis](https://media.springernature.com/m685/springer-static/image/art%3A10.1038%2Fs41597-025-05997-4/MediaObjects/41597_2025_5997_Fig1_HTML.png)
Data collection
This dataset includes WB-FDG-PET/CT imaging data and subsequently derived CT and PET readouts for 135 organs of 48 healthy controls: 25 (52%) females and 23 (48%) males. All subjects were adults with a mean age of 38 ± 14 years (range: 19–59 y/o). The status of health, i.e., “healthy”, was defined as the known absence of systemic diseases, such as cancer or diabetes, as verified during the recruitment and initial subject consultation.
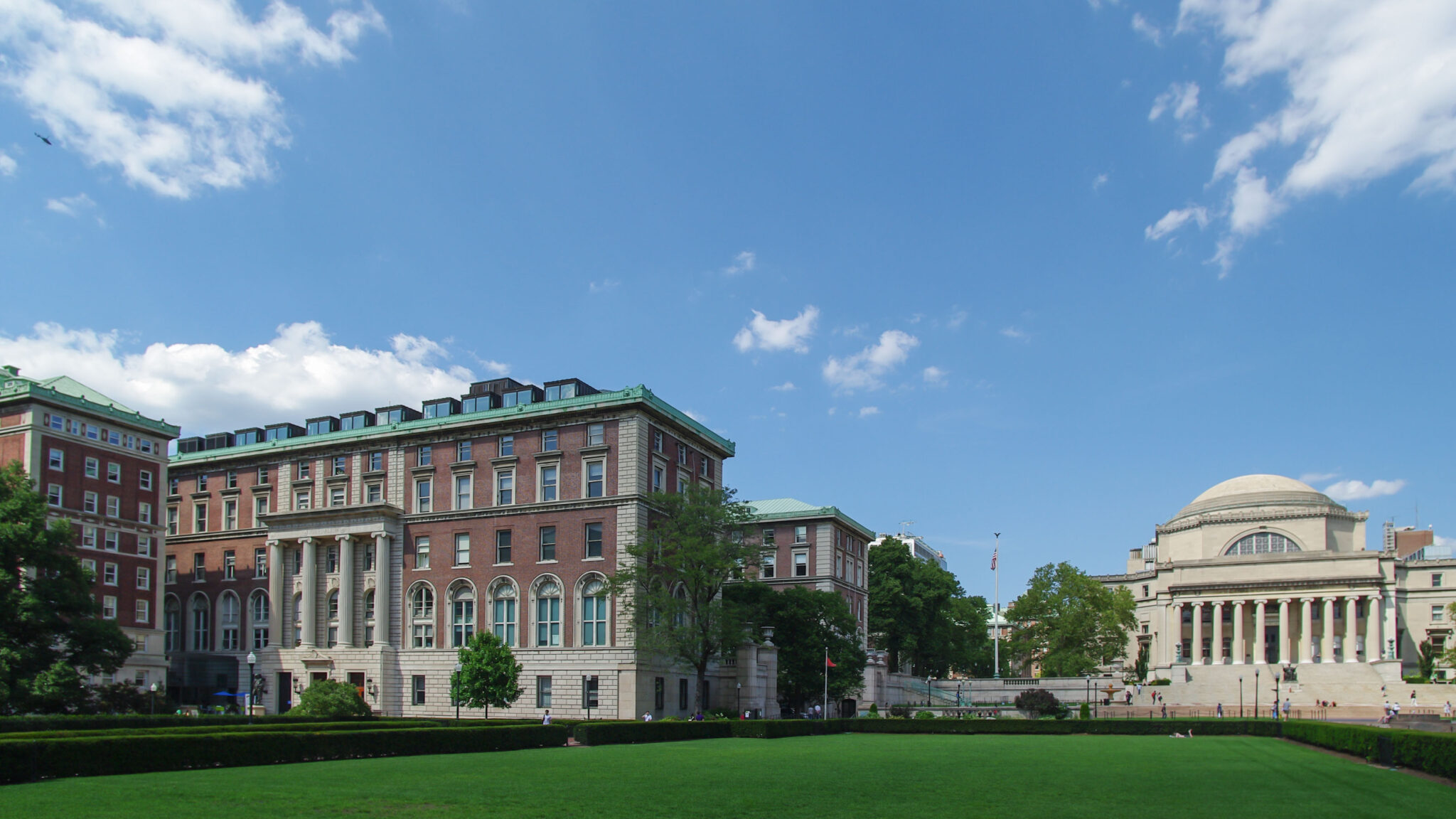
All verified healthy controls were included in a 6-month study (Fig. 1). Clinical tests for stress markers (hair, blood), inflammatory and other biochemical markers (blood) were performed four times throughout the study period: on day 1, the day of the Test PET/CT scan, the day of the Retest PET/CT scan, and again on the last day of the study. The time between the Test and the Retest PET/CT was 4-5 weeks. All subjects were given a wearable device (Garmin Forerunner 955) on day 1 and asked to wear it continuously throughout the entire study period to document biosignals against which any variations in the metabolic patterns seen on the FDG-PET images of the Test and Retest scan will be compared at a later time.

Healthy control study protocol: Recruited subjects were enrolled on day 1 for a clinical assessment, incl. blood parameters and stress markers (“Initiation”). On that day, subjects were given the smartwatch and instructed to wear it continuously throughout the entire study period. After ~5 weeks (day ~35), subjects were invited to the 1st PET/CT scan (Test) and the collection of blood and stress markers. After 4-5 weeks (day ~70), the same procedure (Retest) was repeated, and subjects were released for continuous wearable monitoring. After ~100 days, the subjects were called in for their clinical assessment, incl. blood parameters and stress markers (“Release”). Also, all subjects handed over their smartwatch, and their bio readouts were extracted and anonymized to be stored locally on-site.
All datasets were acquired in accordance with the Declaration of Helsinki, with written informed consent obtained from all subjects prior to examinations. All data were acquired between Sep 28, 2023, and Aug 5, 2024, at the Medical University of Vienna, Austria, following IRB approval (EK 1707/2022), which can also be found in the public registry of approved studies of the IRB: The demographics of the study participants – recruited via the University network and adverts in cooperating institutions – are summarized in Table S1, and an overview is presented in Fig. 2.

Demographics of the 48 healthy controls included in the complete study. The age, sex, BMI, and Δ BMI distributions of the 48 subjects are presented as histograms. For Δ BMI, the measurements derived from the Test scan served as a reference.
Of note, this study was approved for the enrollment of 50 healthy controls. After the completion of the first FDG-PET/CT scan (Test, Fig. 1), 2/50 controls presented with incidental findings indicative of malignant disease. Following the bioptic confirmation of the image-based findings, both subjects were excluded from the study and transferred to standard clinical workup.
Imaging protocol
PET/CT imaging was conducted using a Biograph Vision QUADRA system (Siemens Healthineers, USA) with software version VR20B. This system integrates a 128-slice dual-energy CT alongside a PET component comprising four VISION PET units35. The system offers a 106 cm axial field of view (FOV), with a transverse and axial spatial resolution of 3.3 mm and 3.8 mm, respectively, at a 1 cm FOV offset. It achieves a sensitivity of 180 kcps/MBq with a peak NEC of 276 kcps at 25 kBq/mL. For detailed technical specifications and performance parameters, refer to36,37,38. To ensure the reliability of quantitative measurements, daily quality control procedures are performed for both PET and CT components, along with regular cross-calibration checks, conducted at least every six months and additionally following any changes to the imaging systems or dose calibrators.
Prior to the Test and Retest PET/CT scan, all subjects were asked to fast for 4 hours (blood glucose level < 150 mg/dL). Subjects were positioned head-first in a supine position with arms down and asked to lie still with the permission to breathe normally (shallow) during the duration of the examination (CT and PET for Test and Retest). A low-dose spiral CT scan of the entire imaging range with an automatic tube current modulation set at 25 mA reference and a tube voltage of 140 kVp using the zinc filter and quality setting 3 was performed first. The scanning parameters included a table feed per rotation of 38 mm and a spiral pitch factor of 1.0. Reconstruction was done using the iterative reconstruction algorithm ADMIRE and using the convolutional kernel Br32f and a reconstruction diameter of 780 mm. The reconstructed images had a pixel spacing of 1.523 mm × 1.523 mm, with a slice thickness of 2 mm. The corresponding matrix size was 512 × 512 × 531 voxels.
Following the completion of the CT scan, subjects were moved to the PET imaging position and injected with 100 MBq [18F]FDG. The PET emission scan was set to start immediately, and listmode emission data were acquired for 62 min. At the end of the examination, the subjects were released and asked to void. The total effective dose received by the participating subjects was estimated to be 3.0 mSv, with 33% from the LD-CT and 67% from the emission scan.
PET image data were reconstructed for 57–62 min post-injection using OP-OSEM with TOF information and PSF correction with 4 iterations and 5 subsets. A matrix size of 440 × 440 × 531 voxels with an in-plane voxel spacing of 1.65 mm × 1.65 mm and a slice thickness of 2 mm was used, and a Gaussian post-reconstruction filter of 2 mm full width at half maximum (FWHM) was applied. All standard corrections were utilized, comprising of CT-based attenuation and scatter correction, as well as corrections for randoms, decay, and dead time.
CT and PET reconstruction of all healthy subjects shared in this cohort resulted in 48 PET image volumes matched to 48 CT image sets at two time points along the study trajectory (Fig. 1: Test and Retest). Both PET/CT imaging sessions were set apart by an average of (38 ± 10) days (minimum: 22 days, maximum: 98 days, Table S1). All PET images were converted from Bq/mL to weight-based standardized uptake values (SUV). See Table S1 for details on key protocol parameters and reported scan and subject differences between Test and Retest.
Data processing, image segmentation, and readouts
All 50 participants of this study had their PET/CT examinations reviewed by a board-certified nuclear medicine physician for image quality assessment and incidental findings. As stated, 2/50 enrolled subjects were excluded from this study after confirming their incidental findings on the Test PET/CT (Fig. 1). The acquired PET and CT images of the remaining healthy controls (n = 48) were deemed of sufficiently high quality and free of detrimental artifacts. The images were segmented using the in-house developed and released MOOSE software version 3.0.1325, which was trained on a cohort of 1683 individual cases39, not including the presented cohort. For each subject and dataset (Test, Retest), MOOSE was used to automatically delineate 135 organs from the CT images. Since the CT and PET images were reconstructed with different voxel and matrix sizes, any CT-based multi-organ segmentation mask must be resampled – using a nearest-neighbor interpolation – to match the PET voxel and matrix sizes before extracting complementary regional PET and CT image quantification. Of note, prior to resampling, no additional PET-CT registration was performed for the combined PET/CT data.
In total, 11 biometric parameters were retrieved from both the PET and CT images per organ and subject. These parameters included: six parameters from the CT image (mean, standard deviation (STD), median, minimum, maximum tissue densities in HU, and organ volume), and five corresponding regional values from the matching PET images (mean, STD, median, minimum, and maximum SUV). We evaluated the mean and STD across the 48 subjects at Test and Retest (Fig. 1) as estimates of the normative, cohort-specific organ parameters at both time points. All data processing and metric extraction were done with Python 3.10, utilizing the SimpleITK40,41 package.
While the STD of the normative values across the 135 segmented regions can be estimated from each individual organ (Table S2), we limit our robustness analysis – that is, the Test/Retest variability – to 10 target organs that are known to be involved in a number of prevalent systemic diseases: liver, spleen, lungs (all five lobes), brain, pancreas, kidneys (left and right), heart, subcutaneous fat, visceral fat, and skeletal muscle. Volumes of the latter three tissues were defined at the vertebral L3 level. Here, the L3 level was selected as a well-established surrogate location for global body-composition measurements42,43,44 and more efficient delineation and verification of the fat and muscle segments for subsequent model training.
To evaluate potential sex-based differences in imaging-derived readouts, we performed a group-level comparison using the unpaired Student’s t-test, as a statistical assessment of between-group variability. Further, we assessed the repeatability of the Test and Retest scans at both the group and the subject levels. For the group analysis, we calculated the region-based averages across all 48 subjects, from which we derived statistical differences between Test and Retest readouts using the relative %-difference and the coefficient of variation (CV). We then used a paired Student’s t-test to assess the statistical significance of within-subject repeatability since each subject contributes a directly linked Test-Retest pair. Lastly, the Intraclass Correlation Coefficient (ICC) was used to evaluate repeatability for each organ and parameter45,46, specifically ICC(3,1), which assesses absolute agreement in repeated measures from the same subjects under identical conditions46. For the subject-level analysis, we computed the relative %-difference between the Test and Retest scans for each region on a per-subject basis, averaging these differences across the 48 subjects.
In principle, normative data (Table S2), as included in this healthy cohort, can be used to assess aberrations of tissue metabolism, organ volume, or organ density, as available from cohort-specific readouts of PET (SUV) and CT ([mL], [HU]). Disease-related aberrations can be detected as long as their quantitative impact on CT or PET readouts is larger than the Test/Retest variability. Here, we computed z-scores to highlight subject-specific deviations from the established norm, such as SUV, HU, and volume for each of the 135 segmented organs. The following formula (I) was applied to each organ parameter set to compute the z-score, denoted as \(z\):
$$z=\frac{{mea}{n}_{{subject}}-{mea}{n}_{{cohort}}}{{ST}{D}_{{cohort}}}$$
(I)
The resulting z-scores can be visualized on an organ and voxel level47,48,49. For illustration purposes, we generate an organ atlas that highlights z-scores as deviations from the norm with 2 STDs as a cut-off to filter out noise.
To further expand the usability of this dataset in terms of chronobiological assessments, we report the start time of the Test and Retest PET/CT acquisition for each subject in Table 1. The time of day column indicates whether scans were performed before noon, in the afternoon, or the evening.
Facial anonymization of the image data was performed prior to data sharing. Defacing was applied to the PET and CT images of each subject by first extracting a facial segmentation mask from the CT using the “face” task of TotalSegmentator version 2.10.050. This mask was then applied to both the CT and PET images; for PET, the mask was coregistered using nearest-neighbor interpolation. In both modalities, the masked facial region was downsampled and subsequently upsampled to obscure identifiable features. The pixelated region was reinserted into the corresponding original image using the segmentation mask, effectively replacing the original facial anatomy with an unrecognizable version. All image analysis and quantification presented in this study were performed on the original, non-defaced PET and CT images.
link





:max_bytes(150000):strip_icc()/Health-GettyImages-1413749294-1e0959850fd24395b36963b6b3f2e109.jpg)
